 Thomas P. Sakmar
Thomas P. Sakmar
Head of Laboratory
Laboratory of Molecular Biology and Biochemistry
Rockefeller Research Building 510
The Rockefeller University, Box 187
1230 York Avenue
New York, NY 10065
Telephone: 212-327-8288
Fax: 212-327-7904
sakmar@rockefeller.edu
Published Papers
146).
 Miller, J., Agarwal, A., Devi, L. A., Fontanini, K., Hamilton, J. A., Pin, J. P., Shields, D. C., Spek, C. A.,Sakmar, T. P., Kuliopulos, A. & Hunt, S. W., 3rd.
Miller, J., Agarwal, A., Devi, L. A., Fontanini, K., Hamilton, J. A., Pin, J. P., Shields, D. C., Spek, C. A.,Sakmar, T. P., Kuliopulos, A. & Hunt, S. W., 3rd.
Insider Access: Pepducin Symposium Explores a New Apprach to GPCR Modulation.
Ann N Y Acad Sci 1180:E1–E12 (2009).
145).
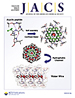 Ahuja, S., Eilers, M., Hirshfeld, A., Yan, E. C. –Y., Ziliox, M., Sakmar, T. P., Sheves, M. & Smith, S. O.
Ahuja, S., Eilers, M., Hirshfeld, A., Yan, E. C. –Y., Ziliox, M., Sakmar, T. P., Sheves, M. & Smith, S. O.
6-s-cis Conformation and Polar Binding Pocket of the Retinal Chromophore in the Photoactivated State of Rhodopsin.
J Am Chem Soc 131:15160–15169 (2009).
144).
 Kapoor, N., Menon, S. T., Chauhan, R., Sachdev, P., & Sakmar, T. P.
Kapoor, N., Menon, S. T., Chauhan, R., Sachdev, P., & Sakmar, T. P.
Structural Evidence for a Sequential Release Mechanism for Activation of Heterotrimeric G Proteins.
J Mol Biol 393:882–897 (2009).
143).
 Ye, S., Huber, T., Vogel, R. & Sakmar, T. P.
Ye, S., Huber, T., Vogel, R. & Sakmar, T. P.
FTIR Analysis of GPCR Activation using Azido Probes
Nat Chem Biol 5:397–399 (2009).
PMID: 19396177 [PubMed - indexed for MEDLINE]
142).
 Ahuja, S., Hornak, V., Yan, E. C. Y., Syrett, N., Goncalves, J. A., Hirshfeld, A., Ziliox, M., Sakmar, T. P., Sheves, M., Reeves, P. J., Smith, S. O. & Eilers, M.
Ahuja, S., Hornak, V., Yan, E. C. Y., Syrett, N., Goncalves, J. A., Hirshfeld, A., Ziliox, M., Sakmar, T. P., Sheves, M., Reeves, P. J., Smith, S. O. & Eilers, M.
Helix Movement Is Coupled to Displacement of the Second Extracellular Loop in Rhodopsin Activation.
Nat Struct Mol Biol 16:168–175 (2009).
MID: 19182802 [PubMed - indexed for MEDLINE]
 141). 141).
Sakmar, T. P. & Huber, T.
Rhodopsin.
in Encyclopedia of Neuroscience. (Squire, L.R., ed.) Oxford: Academic Press. 8:365–372 (2009).
[not indexed for MEDLINE]
140).
 Sakmar, T. P. Sakmar, T. P.
Structure and Function of G Protein-Coupled Receptors: Lessons from Recent Crystal Structures.
in Handbook of Cell Signaling, 2nd edition. (Bradshaw, R. A. & Dennis, E. A., eds.) Oxford: Academic Press. pp. 151–156 (2009).
139).
 Seibert, C., Veldkamp, C.T., Peterson, F.C., Chait, B.T., Volkman, B.F. &Sakmar, T. P.
Seibert, C., Veldkamp, C.T., Peterson, F.C., Chait, B.T., Volkman, B.F. &Sakmar, T. P.
Sequential Tyrosine Sulfation of CXCR4 by Tyrosylprotein Sulfotransferases.
Biochemistry 47:11251–11262 (2008). [*Ranked 19th Most Downloaded Article in Biochemistry Dec. 2008]
PMID: 18834145 [PubMed - indexed for MEDLINE]
138).
 Huber, T., Menon, S. & Sakmar, T. P.
Huber, T., Menon, S. & Sakmar, T. P.
Structural Basis for Ligand Binding and Specificity in Adrenergic Receptors: Implications for GPCR-Targeted Drug Discovery.
Biochemistry 47:11013–11023 (2008).
[*Ranked Most Downloaded Article in Biochemistry Oct. – Dec. 2008; 10th Most Downloaded for past 12 months, Feb. – Mar. 2009; 16th Most Downloaded for past 12 months April – May 2009]
PMID: 18821775 [PubMed - indexed for MEDLINE]
137).
 Veldkamp, C. T., Seibert, C., Peterson, F. C., De la Cruz, N. B., Haugner, III, J. C., Basnet, H., Sakmar, T. P. & Volkman, B. F.
Veldkamp, C. T., Seibert, C., Peterson, F. C., De la Cruz, N. B., Haugner, III, J. C., Basnet, H., Sakmar, T. P. & Volkman, B. F.
Structural Basis of CXCR4 Sulfotyrosine Recognition by the Chemokine SDF-1/CXCL12.
Sci Signal 1:ra4 (2008). [*Cover Article]
PMID: 18799424 [PubMed - indexed for MEDLINE]
136).
 Vogel, R., Mahalingam, M., Lüdeke, S., Huber, T., Siebert, F. & Sakmar, T. P.
Vogel, R., Mahalingam, M., Lüdeke, S., Huber, T., Siebert, F. & Sakmar, T. P.
Functional Role of the “Ionic Lock” – an Interhelical Hydrogen-bond Network in Family A Heptahelical Receptors.
J Mol Biol 380:648–655 (2008).
PMID: 18554610 [PubMed - indexed for MEDLINE]
135).
 Huber, T. & Sakmar, T. P.
Huber, T. & Sakmar, T. P.
Rhodopsin’s Active State Is Frozen Like a DEER in the Headlights.
Proc Natl Acad Sci U S A 105:7343–7344 (2008).
PMID: 18492801 [PubMed - indexed for MEDLINE]
134).
 Banerjee, S., Huber, T. & Sakmar, T. P. Banerjee, S., Huber, T. & Sakmar, T. P.
Rapid Incorporation of Functional Rhodopsin into Nanoscale Apolipoprotein Bound Bilayers (NABB) Particles.
J Mol Biol 377:1067–1081 (2008).
PMID: 18313692 [PubMed - indexed for MEDLINE]
133).
 Louis, M., Huber, T., Benton, R., Sakmar, T. P. & Vosshall, L. B.
Louis, M., Huber, T., Benton, R., Sakmar, T. P. & Vosshall, L. B.
Bilateral Olfactory Senstory Input Enhances Chemotaxis Behavior.
Nat Neurosci 11:187–199 (2008).
PMID: 18157126 [PubMed - indexed for MEDLINE]
132).
 Ye, S., Köhrer, C., Huber, T., Kazmi, M., Sachdev, P., Yan, E. C. Y., Bhagat, A., Rajbhandary, U. L. & Sakmar, T. P.
Ye, S., Köhrer, C., Huber, T., Kazmi, M., Sachdev, P., Yan, E. C. Y., Bhagat, A., Rajbhandary, U. L. & Sakmar, T. P.
Site-specific Incorporation of Keto Amino Acids into Functional G Protein-coupled Receptors Using Unnatural Amino Acid Mutagenesis.
J Biol Chem 283:1525–1533 (2008).
PMID: 17993461 [PubMed - indexed for MEDLINE]
131).
 Seibert, C. & Sakmar, T. P.
Seibert, C. & Sakmar, T. P.
Toward a Framework for Sulfoproteiomics: Synthesis and Characterization of Sulfotyrosine-containing Peptides.
Biopolymers 90:459–477 (2008).
PMID: 17680702 [PubMed - indexed for MEDLINE]
130).
 Janz, J. M., Sakmar, T. P. & Min, K. C.
Janz, J. M., Sakmar, T. P. & Min, K. C.
A Novel Interaction Between AIP4 and bPIX Is Mediated by an SH3 Domain.
J Biol Chem 282:28893–28903 (2007).
PMID: 17652093 [PubMed - indexed for MEDLINE]
129).
 Periole, X., Huber, T., Marrink, S. J. & Sakmar, T. P.
Periole, X., Huber, T., Marrink, S. J. & Sakmar, T. P.
G Protein-coupled Receptors Self-assemble in Dynamics Simulations of Model Bilayers.
J Am Chem Soc 129:10126–10132 (2007).
PMID: 17658882 [PubMed - indexed for MEDLINE]
128).
 Sachdev, P., Menon, S., Kastner, D. B., Chuang, J. Z., Yeh, T. Y., Conde, C., Caceres, A., Sung, C. H. & Sakmar, T. P.
Sachdev, P., Menon, S., Kastner, D. B., Chuang, J. Z., Yeh, T. Y., Conde, C., Caceres, A., Sung, C. H. & Sakmar, T. P.
G Protein bg Sununit Interaction with the Dynein Light-Chain Component Tctex-1 Regulates Neurite Outgrowth.
EMBO J 26:2621–2632 (2007).
PMID: 17491591 [PubMed - indexed for MEDLINE]
127).
 Yan, E. C. Y., Epps., J., Lewis, J. W.Szundi, I., Bhagat, A., Sakmar, T. P. & Kliger, D. S.
Yan, E. C. Y., Epps., J., Lewis, J. W.Szundi, I., Bhagat, A., Sakmar, T. P. & Kliger, D. S.
Photointermediates of the Rhodopsin S186A Mutant As a Probe of the Hydrogen Bond Network in the Chromophore Pocket and the Mechanism of Counterion Switch.
J Phys Chem C 111:8843–8848 (2007).
[not indexed for MEDLINE]
126).
 Vogel, R., Sakmar, T. P., Sheves, M. & Siebert, F.
Vogel, R., Sakmar, T. P., Sheves, M. & Siebert, F.
Coupling of Protonation Switches During Rhodopsin Activation.
Photochem Photobiol 83:286–292 (2007).
PMID: 17576345 [PubMed - indexed for MEDLINE]
125).
 Botelho, A. V., Huber, T., Peterson, F. C., Sakmar, T. P. & Brown, M. F.
Botelho, A. V., Huber, T., Peterson, F. C., Sakmar, T. P. & Brown, M. F.
Curvature and Hydrophobic Forces Drive Oligomerization and Modulate Activity of Rhodopsin in Membranes.
Biophys J 91:4464–4477 (2006).
PMID: 17012328 [PubMed - indexed for MEDLINE]
124).
 Sakmar, T. P.
Sakmar, T. P.
Timing is Everything: Direct Measurement of Retinol Production in Cones and Rods.
J Gen Physiol 128:147–148 (2006).
PMID: 16847095 [PubMed - indexed for MEDLINE]
123).
 Vogel, R., Siebert, F., Yan, E. C. Y., Sakmar, T. P., Hirshfeld, A. & Sheves, M.
Vogel, R., Siebert, F., Yan, E. C. Y., Sakmar, T. P., Hirshfeld, A. & Sheves, M.
Modulating Rhodopsin Receptor Activation by Altering the pKA of the Retinal Schiff Base.
J Am Chem Soc 128:10530–10512 (2006).
PMID: 16895417 [PubMed - indexed for MEDLINE]
122).
 Veldkamp, C. T., Seibert, C., Peterson, F. C., Sakmar, T. P. & Volkman, B. F.
Veldkamp, C. T., Seibert, C., Peterson, F. C., Sakmar, T. P. & Volkman, B. F.
Recognition of a CXCR4 Sulfotyrosine by the Chemokine Stromal Cell-derived Factor-1a (SDF-1a/CXCL12).
J Mol Biol 359:1400–1409 (2006).
PMID: 16725153 [PubMed - indexed for MEDLINE]
121).
 Lewis, J. W., Szundi, I., Kazmi, M. A., Sakmar, T. P. & Kliger, D. S.
Lewis, J. W., Szundi, I., Kazmi, M. A., Sakmar, T. P. & Kliger, D. S.
Proton Movement and Photointermediate Kinetics in Rhodopsin Mutants.
Biochemistry 45:5430–5439 (2006).
PMID: 16634624 [PubMed - indexed for MEDLINE]
120).
 Kazmi, M., Sakmar, T. P. & Yau, K. –W.
Kazmi, M., Sakmar, T. P. & Yau, K. –W.
Parietal-Eye Phototransduction Components: Insight into Vertebrate Photoreceptor Evolution.
Science 311:1617–1621 (2006).
PMID: 16543463 [PubMed - indexed for MEDLINE]
119).
 Hoelz, A., Janz, J. M., Lawrie, S. D., Corwin, B., Lee, A. & Sakmar, T. P.
Hoelz, A., Janz, J. M., Lawrie, S. D., Corwin, B., Lee, A. & Sakmar, T. P.
Crystal Structure of the SH3 Domain of bPIX in Complex with a High Affinity Peptide from PAK2.
J Mol Biol 358:509–522 (2006).
PMID: 16527308 [PubMed - indexed for MEDLINE]
118).
 Seibert, C., Ying, W., Gavrilov, S., Tsamis, F., Kuhmann, S. E., Palani, A., Tagat, J. R., Clader, J. W., McCombie, S. W., Baroudy, B. M., Smith, S. O., Dragic, T., Moore, J. P. & Sakmar, T. P. Seibert, C., Ying, W., Gavrilov, S., Tsamis, F., Kuhmann, S. E., Palani, A., Tagat, J. R., Clader, J. W., McCombie, S. W., Baroudy, B. M., Smith, S. O., Dragic, T., Moore, J. P. & Sakmar, T. P.
Interaction of Small Molecule Inhibitors of HIV-1 Entry with CCR5.
Virology 349:41–54 (2006).
PMID: 16494916 [PubMed - indexed for MEDLINE]
117).
 Vogel, R., Ludeke, S., Siebert, F., Sakmar, T. P., Hirshfeld, A. & Sheves, M.
Vogel, R., Ludeke, S., Siebert, F., Sakmar, T. P., Hirshfeld, A. & Sheves, M.
Agonists and Partial Agonists of Rhodopsin: Retinal Polyene Methylation Affects Receptor Activation.
Biochemistry 45:1640–1652 (2006).
PMID: 16460011 [PubMed - indexed for MEDLINE]
116).
 Ludeke, S., Beck, M., Yan, E. C., Sakmar, T. P., Siebert, F. & Vogel, R.
Ludeke, S., Beck, M., Yan, E. C., Sakmar, T. P., Siebert, F. & Vogel, R.
The Role of Glu181 in the Photoactivation of Rhodopsin.
J Mol Biol 353:345–356 (2005).
PMID: 16169009 [PubMed - indexed for MEDLINE]
115).
 Huber, T. & Sakmar, T. P. Huber, T. & Sakmar, T. P.
The Photoreceptor Membrane As a Model System in the Study of Biological Signal Transduction.
in Advances in Planar Lipid Bilayers and Liposomes. (Tien, H. T. & Ottova, A., eds.) Elsevier, Oxford, pp. 182–206 (2005).
[not indexed for MEDLINE]
114).
 Sakmar, T. P.
Sakmar, T. P.
Twenty Years of the Magnificent Seven.
The Scientist 19:22-23 (2005). [*Cover Article]
113).
 Bartfai, T., Benovic, J. L., Bockaert, J., Bond, R. A., Bouvier, M., Christopoulos, A., Civelli, O., Devi, L. A., George, S. R., Inui, A., Kobilka, B., Leurs, R., Neubig, R., Pin, J. –P., Quirion, R., Roques, B. P., Sakmar, T. P., Seifert, R., Stenkamp, R. E. & Strange, P. G.
Bartfai, T., Benovic, J. L., Bockaert, J., Bond, R. A., Bouvier, M., Christopoulos, A., Civelli, O., Devi, L. A., George, S. R., Inui, A., Kobilka, B., Leurs, R., Neubig, R., Pin, J. –P., Quirion, R., Roques, B. P., Sakmar, T. P., Seifert, R., Stenkamp, R. E. & Strange, P. G.
Twenty Questions: The State of GPCR Research in 2004.
Nat Revs Drug Disc 3:575–626 (2004).
[not indexed for MEDLINE]
112).
 Yan, E. C., Ganim, Z., Kazmi, M. A., Chang, B. S. W., Sakmar, T. P. & Mathies, R. A.
Yan, E. C., Ganim, Z., Kazmi, M. A., Chang, B. S. W., Sakmar, T. P. & Mathies, R. A.
Resonance Raman Analysis of the Mechanism of Energy Storage and Chromophore Distortion in the Primary Visual Photoproduct.
Biochemistry 43:10867–10876 (2004).
PMID: 15323547 [PubMed - indexed for MEDLINE]
111).
 Lewis, J. W., Szundi, I., Kazmi, M. A., Sakmar, T. P. & Kliger, D. S.
Lewis, J. W., Szundi, I., Kazmi, M. A., Sakmar, T. P. & Kliger, D. S.
Time-Resolved Photointermediate Changes in Rhodopsin Glu181 Mutants.
Biochemistry 43:12614–12621 (2004).
PMID: 15449951 [PubMed - indexed for MEDLINE]
110).
 Billick, E., Seibert, C., Pugach, P., Ketas, T., Trkola, A., Endres, M. J., Murgolo, N. J., Coates, E., Reyes, G. R., Baroudy, B. M., Sakmar, T. P., Moore, J. P. & Kuhmann, S. E.
Billick, E., Seibert, C., Pugach, P., Ketas, T., Trkola, A., Endres, M. J., Murgolo, N. J., Coates, E., Reyes, G. R., Baroudy, B. M., Sakmar, T. P., Moore, J. P. & Kuhmann, S. E.
The Differential Sensitivity of Human and Rhesus Macaque CCR5 to Small-Molecule Inhibitors of Human Immunodeficiency Virus Type 1 Entry Is Explained by a Single Amino Acid Difference and Suggests a Mechanism of Action for These Inhibitors.
J Virol 78:4134–4144 (2004).
PMID: 15047829 [PubMed - indexed for MEDLINE]
109).
 Seibert, C., & Sakmar, T. P. Seibert, C., & Sakmar, T. P.
Small-Molecule Antagonists of CCR5 and CXCR4: A Promising New Class of Anti-HIV-1 Drugs.
Curr Pharm Design 10:2041–2062 (2004).
PMID: 15279544 [PubMed - indexed for MEDLINE]
108).
 Sakmar, T. P. Sakmar, T. P.
Rhodopsin.
in Encyclopedia of Neuroscience. 3rd Ed. (Adelman, G. & Smith, B. H., eds.) Elsevier Science B.V., Amsterdam, Holland (2004).
[not indexed for MEDLINE]
107).
 Sakmar, T. P. Sakmar, T. P.
Structure and Function of G Protein-Coupled Receptors: Lessons from the Crystal Structure of Rhodopsin.
in Handbook of Cell Signaling, Academic Press, San Diego, CA, pp. 139–143 (2003).
[not indexed for MEDLINE]
106).
 Yan, E. C., Kazmi, M. A., Ganim, Z., Hou, J. M., Pan, D., Chang, B. S. W., Sakmar, T. P. & Mathies, R. A.
Yan, E. C., Kazmi, M. A., Ganim, Z., Hou, J. M., Pan, D., Chang, B. S. W., Sakmar, T. P. & Mathies, R. A.
Retinal Counterion Switch in the Photoactivation of the G Protein-Coupled Receptor Rhodopsin.
Proc Natl Acad Sci U S A 100:9262–9267 (2003).
[*See commentary: Birge, R. R. & Knox, B. E. Perspectives on the Counterion Switch-Induced Photoactivation of the G Protein-Coupled Receptor Rhodopsin. Proc Natl Acad Sci U S A 100:9105–9107 (2003).]
PMID: 12835420 [PubMed - indexed for MEDLINE]
105).
 Tsamis, F., Gavrilov, S., Kajumo, F., Seibert, C., Kuhmann, S., Ketas, T., Trkola, A., Palani, A., Clader, J. W., Tagat, J. R., McCombie, S., Baroudy,B., Moore, J. P., Sakmar, T. P. & Dragic, T.
Tsamis, F., Gavrilov, S., Kajumo, F., Seibert, C., Kuhmann, S., Ketas, T., Trkola, A., Palani, A., Clader, J. W., Tagat, J. R., McCombie, S., Baroudy,B., Moore, J. P., Sakmar, T. P. & Dragic, T.
Analysis of the Mechanism by Which the Small-Molecule CCR5 Antagonists SCH-351125 and SCH-350581 Inhibit Human Immunodeficiency Virus Type 1 Entry.
J Virol 77:5201–5208 (2003).
PMID: 12692222 [PubMed - indexed for MEDLINE]
104).
 Sakmar, T. P.
Sakmar, T. P.
Color Vision.
in Adler's Physiology of the Eye, 10th Edition. (Kaufmann, P. L. & Alm, A., eds.)
Mosby, St. Louis, pp. 578–585 (2003).
[not indexed for MEDLINE]
103).
 Seibert, C., Cadene, M., Sanfiz, A., Chait, B. T. & Sakmar, T. P.
Seibert, C., Cadene, M., Sanfiz, A., Chait, B. T. & Sakmar, T. P.
Tyrosine Sulfation of CCR5 N-Terminal Peptide by Tyrosylprotein Sulfotransferases 1 and 2 Follows a Discrete Pattern and Temporal Sequence.
Proc Natl Acad Sci U S A 99:11031–11036 (2002).
PMID: 12169668 [PubMed - indexed for MEDLINE]
102).
 Unson, C. G., Wu, C. –R., Yoo, B., Cheung, C., Jiang, Y., Sakmar, T. P. & Merrifield, R. B.
Unson, C. G., Wu, C. –R., Yoo, B., Cheung, C., Jiang, Y., Sakmar, T. P. & Merrifield, R. B.
The Roles of Specific Extracellular Domains of the Glucagon Receptor in Ligand Binding and Signaling.
Biochemistry 41:11795–11803 (2002).
PMID: 12269822 [PubMed - indexed for MEDLINE]
101).
 De, S. & Sakmar, T. P.
De, S. & Sakmar, T. P.
Interaction of A2E with Model Membranes. Implications to the Pathogenesis of Age-Related Macular Degeneration.
J Gen Physiol 120:147–157 (2002).
PMID: 12149277 [PubMed - indexed for MEDLINE]
100).
 Gopala Krishna, A., Menon, S. T., Terry, T. J.& Sakmar, T. P. Gopala Krishna, A., Menon, S. T., Terry, T. J.& Sakmar, T. P.
Evidence That Helix 8 of Rhodopsin Acts As a Membrane-Dependent Conformational Switch.
Biochemistry 41:8298–8309 (2002).
PMID: 12081478 [PubMed - indexed for MEDLINE]
99).
Chang, B. S. W., Jönsson, K., Kazmi, M. A., Donoghue, M. J. & Sakmar, T. P.
Recreating a Functional Ancestral Archosaur Visual Pigment.
Mol Biol Evol 19:1483–1489 (2002). [*Rated “EXCEPTIONAL” by the Faculty of 1000]
PMID: 12200476 [PubMed - indexed for MEDLINE]
98).
 Marin, E. P., Gopala Krishna, A. & Sakmar, T. P.
Marin, E. P., Gopala Krishna, A. & Sakmar, T. P.
Disruption of the a5 Helix of Transducin Impairs Rhodopsin-Catalyzed Nucleotide Exchange.
Biochemistry 41:6988–6994 (2002).
PMID: 12033931 [PubMed - indexed for MEDLINE]
97).
 Yan, E. C. Y., Kazmi, M. A., De, S., Chang, B. S. W., Seibert, C., Marin, E. P., Mathies, R. A. & Sakmar, T. P. Yan, E. C. Y., Kazmi, M. A., De, S., Chang, B. S. W., Seibert, C., Marin, E. P., Mathies, R. A. & Sakmar, T. P.
Function of Extracellular Loop 2 in Rhodopsin: Glutamic Acid 181 Modulates Stability and Absorption Wavelength of Metarhodopsin II.
Biochemistry 41:3620–3627 (2002).
PMID: 11888278 [PubMed - indexed for MEDLINE]
96).
Sakmar, T. P.
The Structure of Rhodopsin and the Superfamily of Seven-Helical Receptors: The Same and Not the Same.
Curr Opin Cell Biol 14:189–195 (2002).
PMID: 11891118 [PubMed - indexed for MEDLINE]
95).
 Sakmar, T. P., Menon, S. T., Marin, E. P. & Awad, E. S. Sakmar, T. P., Menon, S. T., Marin, E. P. & Awad, E. S.
Rhodopsin: Insights from Recent Structural and Mutagenesis Studies.
Annu Rev Biophys Biomol Struct 31:443–484 (2002).
PMID: 11988478 [PubMed - indexed for MEDLINE]
94).
 Chang, B. S. W., Kazmi, M. & Sakmar, T. P. Chang, B. S. W., Kazmi, M. & Sakmar, T. P.
Synthetic Gene Technology: Applications to Ancestral Gene Reconstruction and Structure-Function Studies of Receptors.
Methods Enzymol 343:274–294 (2002).
PMID: 11665573 [PubMed - indexed for MEDLINE]
93).
 Jiang, Y., Cypess, A. M., Muse, E. D., Wu, C-. R., Unson, C. G., Merrifield, R. B. & Sakmar, T. P. Jiang, Y., Cypess, A. M., Muse, E. D., Wu, C-. R., Unson, C. G., Merrifield, R. B. & Sakmar, T. P.
Glucagon Receptor Activates Extracellular Signal-Regulated Protein Kinase (ERK) 1/2 via cAMP-Dependent Protein Kinase.
Proc Natl Acad Sci U S A 98:10102–10107 (2001).
PMID: 11517300 [PubMed - indexed for MEDLINE]
92).
 Menon, S. T., Han, M. & Sakmar, T. P.
Menon, S. T., Han, M. & Sakmar, T. P.
Rhodopsin: Structural Basis of Molecular Physiology.
Physiol Revs 81:1659–1688 (2001). [*Cover Article]
PMID: 11581499 [PubMed - indexed for MEDLINE]
91).
 Marin, E. P., Krishna, A. G. & Sakmar, T. P. Marin, E. P., Krishna, A. G. & Sakmar, T. P.
Rapid Activation of Transducin by Mutations Distant From the Nucleotide-Binding Site: Evidence for a Mechanistic Model of Receptor-Catalyzed Nucleotide Exchange by G Proteins.
J Biol Chem 276:27400–27405 (2001).
PMID: 11356823 [PubMed - indexed for MEDLINE]
90).
 Marin, E. P., Krishna, A. G., Archambault, V., Simuni, E., Fu, W. Y. & Sakmar, T. P.
Marin, E. P., Krishna, A. G., Archambault, V., Simuni, E., Fu, W. Y. & Sakmar, T. P.
The Function of Interdomain Interactions in Controlling Nucleotide Exchange Rates in Transducin.
J Biol Chem 276:23873–23880 (2001).
PMID: 11290746 [PubMed - indexed for MEDLINE]
89).
 Cypess, A. M., Muse, E. D., Wu, C-. R., Unson, C. G. & Sakmar, T. P.
Cypess, A. M., Muse, E. D., Wu, C-. R., Unson, C. G. & Sakmar, T. P.
Glucagon Receptor Causes Glucagon-Dependent Activation of Erk 1/2 in H22 Stable Cell Lines.
in Peptides for the New Millennium. (Fields, G. B., Barany, G. & Tam, J. P., eds.) Kluwer Academic Press, Dordrecht, The Netherlands, pp. 600–601 (2000).
[not indexed for MEDLINE]
88).
 Sakmar, T. P. Sakmar, T. P.
Restoration of Compact Discs.
Nature Genet 25:245–246 (2000).
PMID: 10888860 [PubMed - indexed for MEDLINE]
87).
 Lin, S. W., Han, M., & Sakmar, T. P. Lin, S. W., Han, M., & Sakmar, T. P.
Analysis of Functional Microdomains of Rhodopsin.
Methods Enzymol 315:116–130 (2000).
PMID: 10736698 [PubMed - indexed for MEDLINE]
86).
 Fahmy, K., Sakmar, T. P., & Siebert, F.
Fahmy, K., Sakmar, T. P., & Siebert, F.
Structural Determinants of Active Conformation of Rhodopsin: Molecular Biophysics Approaches.
Methods Enzymol 315:178–196(2000).
PMID: 10736702 [PubMed - indexed for MEDLINE]
85).
 Han, M. & Sakmar, T. P. Han, M. & Sakmar, T. P.
Assays for the Activation of Recombinant Expressed Opsin by all-trans-Retinals.
Methods Enzymol 315:251–267 (2000).
PMID: 10736707 [PubMed - indexed for MEDLINE]
84).
 Isele, J., ,Sakmar T. P. & Siebert, F.
Isele, J., ,Sakmar T. P. & Siebert, F.
Rhodopsin Activation Affects the Environment of Specific Neighboring Phospholipids: An FTIR Spectroscopic Study.
Biophys J 79:3063–3071 (2000).
PMID: 11106612 [PubMed - indexed for MEDLINE]
83).
Min, K. C., Gravina, S. A., & Sakmar, T. P.
Reconstitution of the Vertebrate Visual Cascade Using Recombinant Transducin Purified from Sf9 Cells.
Protein Expr Purification 20:514–526 (2000).
PMID: 11087692 [PubMed - indexed for MEDLINE]
82).
Fahmy, K., Sakmar, T. P. & Siebert, F.
Transducin-dependent Protonation of Glutamic Acid 134 in Rhodopsin.
Biochemistry 39:10607–10612 (2000).
PMID: 10956053 [PubMed - indexed for MEDLINE]
81).
 Unson, C. G., Wu, C. –R., Sakmar, T. P.& Merrifield, R. B.
Unson, C. G., Wu, C. –R., Sakmar, T. P.& Merrifield, R. B.
Selective Stabilization of the High-affinity Binding Conformation of Glucagon Receptor by the Long Splice Variant of Gas.
J Biol Chem 275:21631–21638 (2000).
PMID: 10791965 [PubMed - indexed for MEDLINE]
80).
 Cormier, E. G., Persuh, M., Thompson, D. A. D., Lin, S. W., Sakmar, T. P., Olson, W. C. & Dragic, T.
Cormier, E. G., Persuh, M., Thompson, D. A. D., Lin, S. W., Sakmar, T. P., Olson, W. C. & Dragic, T.
Specific Interaction of CCR5 Amino-terminal Domain Peptides Containing Sulfo-Tyrosines with HIV-1 Envelope Glycoprotein gp120.
Proc Natl Acad Sci U S A 97:5762–5767 (2000).
PMID: 10823934 [PubMed - indexed for MEDLINE]
79).
 Dragic, T., Trkola, A., Thompson, D. A. D., Cormier, E. G., Kajumo, F. A., Maxwell, E., Lin, S. W., Ying, W., Smith, S. O., Sakmar, T. P. & Moore, J. P.
Dragic, T., Trkola, A., Thompson, D. A. D., Cormier, E. G., Kajumo, F. A., Maxwell, E., Lin, S. W., Ying, W., Smith, S. O., Sakmar, T. P. & Moore, J. P.
A Binding Pocket for a Small Molecule Inhibitor of HIV-1 Entry Within the Transmembrane Helices of CCR5.
Proc Natl Acad Sci U S A 97:5639–5644 (2000).
PMID: 10779565 [PubMed - indexed for MEDLINE]
78).
Kazmi, M., Snyder, L. A., Cypess, A. M., Graber, S. G. & Sakmar, T. P.
Selective Reconstitution of Human D4 Dopamine Receptor Variants with Gia Subtypes.
Biochemistry 38:3734–3744 (2000).
PMID: 10736173 [PubMed - indexed for MEDLINE]
77).
 Ernst, O. E., Meyer, C., Marin, E. P., Fu, W. –Y., Sakmar, T. P. & Hofmann, K. P.
Ernst, O. E., Meyer, C., Marin, E. P., Fu, W. –Y., Sakmar, T. P. & Hofmann, K. P.
Mutation of the Fourth Cytoplasmic Loop of Rhodopsin Affects Binding of Transducin and Peptides Derived from the Carboxyl-terminal Sequences of Transducin a and g Subunits.
J Biol Chem 275:1937–1943 (2000).
PMID: 10636895 [PubMed - indexed for MEDLINE]
76).
 Marin, E. P., Gopala Krishna, A., Zyvaga, T. A., Isele, J., Siebert, F. & Sakmar, T. P. Marin, E. P., Gopala Krishna, A., Zyvaga, T. A., Isele, J., Siebert, F. & Sakmar, T. P.
The Amino-terminal Region of the Fourth Cytoplasmic Loop of Rhodopsin Modulates Rhodopsin-transducin Interaction.
J Biol Chem 275:1930–1936 (2000).
PMID: 10636894 [PubMed - indexed for MEDLINE]
75).
Lewis, J. W., Szundi, I., Fu, W. -Y., Sakmar, T. P. & Kliger, D. S.
pH Dependence of Photolysis Intermediates in the Photoactivation of Rhodopsin Mutant E113Q.
Biochemistry 39:599–606 (2000).
PMID: 10642185 [PubMed - indexed for MEDLINE]
74).
 Sakmar, T. P.
Sakmar, T. P.
Rhodopsin Early Receptor Potential Revisited.
Biophys J 77:1189–1190 (1999).
PMID: 10465733 [PubMed - indexed for MEDLINE]
PMCID: PMC1300410
73).
 Lin, S. W. & Sakmar, T. P. Lin, S. W. & Sakmar, T. P.
Colour Tuning Mechanism of Visual Pigments.
in Rhodopsins and Photo-transduction. Novartis Foundation Symposium No. 224. (Goode, J., ed.) John Wiley & Sons, Inc., New York, vol. 224, pp. 124–141 (1999).
PMID: 10614049 [PubMed - indexed for MEDLINE]
72).
 Sakmar, T. P.
Sakmar, T. P.
Rhodopsin.
in The Encyclopedia of Neuroscience. (Adelman, G. & Smith, B. H., eds.) Elsevier Science B.V., Amsterdam, Holland, pp. 1807–1810 (1999).
[not indexed for MEDLINE]
71).
 Sakmar, T. P. & Min, K. C. Sakmar, T. P. & Min, K. C.
Lessons from Rhodopsin.
in Receptor Biochemistry and Methodology. Vol III - Structure/Function Analysis of GPCRs. (Wess, J., ed.) John Wiley & Sons, Inc., New York, vol. 3, pp. 85–107 (1999).
[not indexed for MEDLINE]
70).
 Kochendoerfer, G. G., Lin, S. W., Sakmar, T. P. & Mathies, R. A.
Kochendoerfer, G. G., Lin, S. W., Sakmar, T. P. & Mathies, R. A.
How Color Visual Pigments Are Tuned.
Trends Biochem Sci 24:300–305 (1999).
PMID: 10431173 [PubMed - indexed for MEDLINE]
69).
 Simpson, M. M., Ballesteros, J. A., Chiappa, V., Chen, J., Suehiro, M., Hartman, D. S., Godel, T., Snyder, L. A., Sakmar, T. P. & Javitch, J. A.
Simpson, M. M., Ballesteros, J. A., Chiappa, V., Chen, J., Suehiro, M., Hartman, D. S., Godel, T., Snyder, L. A., Sakmar, T. P. & Javitch, J. A.
Dopamine D4/D2 Receptor Selectivity Is Determined by a Divergent Aromatic Microdomain Contained within the Second, Third, and Seventh Membrane-Spanning Segments.
Mol Pharmacol 56:1116–1126 (1999).
PMID: 10570038 [PubMed - indexed for MEDLINE]
68).
Cypess, A. M., Unson, C. G., Wu, C. –R. & Sakmar, T. P.
Two Cytoplasmic Loops of the Glucagon Receptor Are Required to Elevate cAMP or Intracellular Calcium.
J Biol Chem 274:19455–19464 (1999).
PMID: 10383462 [PubMed - indexed for MEDLINE]
67).
 Sakmar, T. P.
Sakmar, T. P.
Rhodopsin: a Prototypical G Protein-coupled Receptor.
Progr Nucleic Acid Res Mol Biol 59:1–34 (1998).
PMID: 9427838 [PubMed - indexed for MEDLINE]
66).
 Beck, M., Siebert, F. & Sakmar, T. P.
Beck, M., Siebert, F. & Sakmar, T. P.
Evidence for the Specific Interaction of a Lipid Molecule with Rhodopsin Which Is Altered in the Transition to the Active State Metarhodopsin II.
FEBS Lett 436:304–308 (1998).
PMID: 9801137 [PubMed - indexed for MEDLINE]
65).
Lin, S. W., Kochendoerfer, G. G., Carroll, K. S., Wang, D., Mathies, R. A. & Sakmar, T. P.
Mechanisms of Spectral Tuning in Blue Cone Visual Pigments. Visible and Raman Spectroscopy of Blue-Shifted Rhodopsin Mutants.
J Biol Chem 273:24583–24591 (1998).
PMID: 9733753 [PubMed - indexed for MEDLINE]
64).
Beck, M., Sakmar, T. P. & Siebert, F.
Spectroscopic Evidence for Interaction Between Transmembrane Helices 3 and 5 in Rhodopsin.
Biochemistry 37:7630–7639 (1998).
PMID: 9585578 [PubMed - indexed for MEDLINE]
63).
Han, M., Smith, S. O. & Sakmar, T. P.
Constitutive Activation of Opsin by Mutation of Methionine 257 on Transmembrane Helix 6.
Biochemistry 37:8253–8261 (1998).
PMID: 9609722 [PubMed - indexed for MEDLINE]
62).
 Dragic, T., Trkola, A., Lin, S. W., Nagashima, K., Kajumo, F., Allaway, G., Wu, L., MacKay, C., Sakmar, T. P., Maddon, P. J. & Moore, J. P.
Dragic, T., Trkola, A., Lin, S. W., Nagashima, K., Kajumo, F., Allaway, G., Wu, L., MacKay, C., Sakmar, T. P., Maddon, P. J. & Moore, J. P.
N-Terminal Substitutions in the CCR5 Co-receptor Impair gp120 Binding and HIV-1 Entry.
J Virol 72:279–284 (1998).
PMID: 9420225 [PubMed - indexed for MEDLINE]
61).
Han, M., Groesbeek, M., Smith, S. O. & Sakmar, T. P.
The Role of the C9-Methyl Group in Rhodopsin Activation: Characterization of Mutant Opsins with the Artificial Chromophore 11-cis-9-demethyl-Retinal.
Biochemistry 37:538–545 (1998).
PMID: 9425074 [PubMed - indexed for MEDLINE]
60).
Donzella, G. A., Schols, D., Lin, S. W., Esté, J. A., Nagashima, K. A., Maddon, P. J., Allaway, G. P., Sakmar, T. P., Henson, G., De Clercq, E. & Moore, J. P.
AMD3100, a Small Molecule Inhibitor of HIV-1 Entry via the CXCR4 Co-receptor.
Nature Med 4:72–77 (1998).
PMID: 9427609 [PubMed - indexed for MEDLINE]
59).
 Sakmar, T. P. Sakmar, T. P.
Chemistry of Vision.
in Macmillan Encyclopedia of Chemistry . (Lagowski, J. J., ed.) Macmillan Reference USA, New York, vol. 4, pp. 1492–1495 (1997).
[not indexed for MEDLINE]
58).
 Sakmar, T. P. Sakmar, T. P.
Rhodopsin.
in The Encyclopedia of Neuroscience . (Adelman, G. & Smith, B. H., eds.) Elsevier Science B.V., Amsterdam, Holland, CD-ROM (1997).
[not indexed for MEDLINE]
57).
Han, M., Groesbeek, M., Sakmar, T. P. & Smith, S. O.
The C9-Methyl Group of Retinal Interacts with Glycine 121 in Rhodopsin.
Proc Natl Acad Sci U S A 94:13442–13447 (1997).
PMID: 9391044 [PubMed - indexed for MEDLINE]
56).
Jäger, S., Han, M., Lewis, J. W., Szundi, I., Sakmar, T. P. & Kliger, D. S.
Properties of Early Photolysis Intermediates of Rhodopsin Are Regulated by Glycine 121 and Phenylalanine 261.
Biochemistry 36:11804–11810 (1997).
PMID: 9305971 [PubMed - indexed for MEDLINE]
55).
Han, M., Lou, J., Nakanishi, K., Sakmar, T. P. & Smith, S. O.
Partial Agonist Activity of 11-cis-Retinal in Rhodopsin Mutants.
J Biol Chem 272:23081–23085 (1997).
PMID: 9287308 [PubMed - indexed for MEDLINE]
54).
Jäger, S., Lewis, J. W., Zyvaga, T. A., Szundi, I., Sakmar, T. P. & Kliger, D. S.
Chromophore Structural Changes in Rhodopsin From Nanoseconds to Microseconds Following Pigment Photolysis.
Proc Natl Acad Sci U S A 94:8557–8562 (1997).
PMID: 9238015 [PubMed - indexed for MEDLINE]
Sheih, T., Han, M., Sakmar, T. P. & Smith, S. O.
The Steric Trigger in Rhodopsin Activation.
J Mol Biol 269:373–384 (1997).
PMID: 9199406 [PubMed - indexed for MEDLINE]
52).
Jäger, S., Lewis, J. W., Zyvaga, T. A., Szundi, I., Sakmar, T. P. & Kliger, D. S.
Time-resolved Spectroscopy of the Early Photolysis Intermediates of Rhodopsin Schiff Base Counterion Mutants.
Biochemistry 36:1999–2009 (1997).
PMID: 9047297 [PubMed - indexed for MEDLINE]
51).
 Kazmi, M. A., Sakmar, T. P. & Ostrer, H.
Kazmi, M. A., Sakmar, T. P. & Ostrer, H.
Mutation of a Conserved Cysteine in the X-linked Cone Opsins Disrupts Targeting and Causes Color Vision Deficiencies.
Invest Ophthalmol Vis Sci 38:1074–1081 (1997). [*Cover Article]
PMID: 9152227 [PubMed - indexed for MEDLINE]
50).
Sakmar, T. P.
Molecular Mechanism of Signal Transduction by Rhodopsin.
in G Protein-coupled Receptors : New Opportunities for Commercial Development . (Mulford, N., ed.) I.B.C., Inc., Southborough, MA, pp. 638–665 (1996).
49).
Fahmy, K., Zyvaga, T. A., Sakmar, T. P. & Siebert, F.
Spectroscopic Evidence for Altered Chromophore-Protein Interactions in Low Temperature Photoproducts of the Visual Pigment Responsible for Congenital Night Blindness.
Biochemistry 35:15065–15073 (1996).
MID: 8942673 [PubMed - indexed for MEDLINE]
48).
Han, M., Lin, S. W., Minkova, M., Smith, S. O. & Sakmar, T. P.
Functional Interaction of Transmembrane Helices 3 and 6 in Rhodopsin. Replacement of Phenylalanine 261 by Alanine Causes Reversion of Phenotypes of Glycine 121 Replacement Mutants.
J Biol Chem 271:32337–32342 (1996).
PMID: 8943296 [PubMed - indexed for MEDLINE]
47).
Han, M., Lin, S. W., Smith, S. O. & Sakmar, T. P.
The Effects of Amino Acid Replacements of Glycine 121 on Transmembrane Helix 3 of Rhodopsin.
J Biol Chem 271:32330–32336 (1996).
PMID: 8943295 [PubMed - indexed for MEDLINE]
46).
Sheikh, S., Zyvaga, T. A., Lichtarge, O., Sakmar, T. P. & Bourne, H. R.
Rhodopsin Activation Blocked by Metal-Ion-Binding Sites Linking Transmembrane Helices C and F.
Nature 383:347–350 (1996).
PMID: 8848049 [PubMed - indexed for MEDLINE]
45).
Lin, S. W. & Sakmar, T. P.
Specific Tryptophan UV-Absorbance Changes Are Probes of the Transition of Rhodopsin to Its Active State.
Biochemistry 35:11149–11159 (1996).
PMID: 8780519 [PubMed - indexed for MEDLINE]
44).
Zyvaga, T. A., Fahmy, K., Siebert, F. & Sakmar, T. P.
Characterization of the Mutant Visual Pigment Responsible for Congenital Night Blindness: a Biochemical and Fourier-Transform Infrared Spectroscopy Study.
Biochemistry 35:7536–7545 (1996).
PMID: 8652533 [PubMed - indexed for MEDLINE]
43).
Unson, C. G., Cypess, A. M., Wu, C. -R., Goldsmith, P. K., Merrifield, R. B. & Sakmar, T. P.
Antibodies Against Specific Extracellular Epitopes of the Glucagon Receptor Block Glucagon Binding.
Proc Natl Acad Sci U S A 93:310–315 (1996).
PMID: 8552628 [PubMed - indexed for MEDLINE]
42).
Sakmar, T. P. & Fahmy, K.
Properties and Photoactivity of Rhodopsin Mutants.
Israel J Chem 35:325–338 (1995).
[not indexed for MEDLINE]
41).
 Beck, M., Sakmar, T. P. & Siebert, F.
Beck, M., Sakmar, T. P. & Siebert, F.
FTIR Investigations of Rhodopsin and Rhodopsin Mutants in Detergent and Reconstituted into Membranes.
in Sixth European Conference on the Spectroscopy of Biological Molecules (Merlin, J. C., Turrell, S. & Huvenne, J. P., eds.) Kluwer Academic Publishers, Dordrecht, Netherlands, pp. 173–174 (1995).
[not indexed for MEDLINE]
40).
Fahmy, K., Siebert, F. & Sakmar, T. P.
Molecular Determinants of the Active Conformation of Rhodopsin Studied by Attenuated Total Reflectance FTIR Difference Spectroscopy.
in Sixth European Conference on the Spectroscopy of Biological Molecules (Merlin, J. C., Turrell, S. & Huvenne, J. P., eds.) Kluwer Academic Publishers, Dordrecht, Netherlands, pp. 171–172 (1995).
[not indexed for MEDLINE]
39).
Unson, C. G., Cypess, A. M., Kim, H. N., Carruthers, C. J. L., Goldsmith, P. K., Merrifield, R. B. & Sakmar, T. P.
Characterization of Deletion and Truncation Mutants of the Rat Glucagon Receptor: Seven Transmembrane Segments Are Necessary for Receptor Transport to the Plasma Membrane and Glucagon Binding.
J Biol Chem 270:27720–27727 (1995).
PMID: 7499239 [PubMed - indexed for MEDLINE]
38).
 Garcia, P. D., Onrust, R., Bell, S. M., Sakmar, T. P. & Bourne, H. R.
Garcia, P. D., Onrust, R., Bell, S. M., Sakmar, T. P. & Bourne, H. R.
Transducin-a Carboxyl-Terminal Mutations Prevent Activation by Rhodopsin: A New Assay Using Recombinant Proteins Expressed in Cultured Cells.
EMBO J 14:4460–4469 (1995).
PMID: 7556089 [PubMed - indexed for MEDLINE]
37).
 Fahmy, K., Siebert, F. & Sakmar, T. P. Fahmy, K., Siebert, F. & Sakmar, T. P.
The Photoactivated State of Rhodopsin and How It Can Form.
Biophys Chem 56:171–181 (1995).
PMID: 7662864 [PubMed - indexed for MEDLINE]
36).
Ernst, O., Hofmann, K. P. & Sakmar, T. P.
Characterization of Rhodopsin Mutants That Bind Transducin But Fail To Induce GTP Nucleotide Uptake: Classification of Mutant Pigments by Fluorescence, Nucleotide Uptake, and Light-Scattering Assays.
J Biol Chem 270:10580–10586 (1995).
PMID: 7737995 [PubMed - indexed for MEDLINE]
35).
Carruthers, C. J. L. & Sakmar, T. P.
Synthesis and Expression of Synthetic Genes: Applications to Structure-Function Studies of Receptors.
Methods Neurosci 25:322–344 (1995).
[not indexed for MEDLINE]
34).
 Sakmar, T. P. Sakmar, T. P.
Opsins.
in Handbook of Receptors and Channels (Vol. I: G Protein-coupled Receptors) (Peroutka, S. J., ed.) CRC Press, Boca Raton, v. I, pp. 257–276 (1994).
[not indexed for MEDLINE]
33).
Fahmy, K., Siebert, F. & Sakmar, T. P.
A Mutant Rhodopsin Photoproduct with a Protonated Schiff Base Displays an Active-state Conformation: a Fourier-Transform Infrared Spectroscopy Study.
Biochemistry 33:13700–13705 (1994).
PMID: 7947779 [PubMed - indexed for MEDLINE]
32).
Carruthers, C. J. L., Unson, C. G., Kim, H. N. & Sakmar, T. P.
Synthesis and Expression of a Gene for the Rat Glucagon Receptor. Replacement of an Aspartic Acid in the Extracellular Domain Prevents Glucagon Binding.
J Biol Chem 269:29321–29328 (1994).
PMID: 7961903 [PubMed - indexed for MEDLINE]
31).
Jäger, F., Fahmy, K., Sakmar, T. P. & Siebert, F.
Identification of Glutamic Acid 113 as the Schiff Base Proton Acceptor in the Metarhodopsin II Photointermediate of Rhodopsin.
Biochemistry 33:10878–10882 (1994).
PMID: 7916209 [PubMed - indexed for MEDLINE]
30).
Arnis, S., Fahmy, K., Hofmann, K. P. & Sakmar, T. P.
A Conserved Carboxylic Acid Group Mediates Light-Dependent Proton Uptake and Signaling by Rhodopsin.
J Biol Chem 269:23879–23881 (1994).
PMID: 7929034 [PubMed - indexed for MEDLINE]
29).
Zyvaga, T. A., Fahmy, K. & Sakmar, T. P.
Characterization of Rhodopsin-Transducin Interaction: A Mutant Rhodopsin Photoproduct with a Protonated Schiff Base Activates Transducin.
Biochemistry 33:9753-9761 (1994).
PMID: 8068654 [PubMed - indexed for MEDLINE]
28).

Jäger, F., Sakmar, T. P. & Siebert, F.
Molecular Changes of the Membrane-Embedded Carboxyl Group Glu-122 of Bovine Rhodopsin During the Transition to the Active State Metarhodopsin II: An Investigation of the Glu-122 to Asp Mutant Using FT-IR Difference Spectroscopy.
in Fifth International Conference on the Spectroscopy of Biological Molecules (Theophanides, T., Anatassopoulou, J. & Fotopoulos, N., eds.) Kluwer Academic Publishers, Dordrecht, Netherlands, pp. 223-226 (1993).
[not indexed for MEDLINE]
27).
Fahmy, K., Jäger, F., Beck, M., Zyvaga, T. A., Sakmar, T. P. & Siebert, F.
Protonation States of Membrane-Embedded Carboxylic Acid Groups in Rhodopsin and Metarhodopsin II: A Fourier-Transform Infrared Spectroscopy Study of Site-Directed Mutants.
Proc Natl Acad Sci U S A 90:10206–10210 (1993).
PMID: 7901852 [PubMed - indexed for MEDLINE]
26).
Fahmy, K. & Sakmar, T. P.
Light-Dependent Transducin Activation by an Ultraviolet-Absorbing Rhodopsin Mutant.
Biochemistry 32:9165–9171 (1993).
PMID: 8396426 [PubMed - indexed for MEDLINE]
25).
Fahmy, K. & Sakmar, T. P.
Regulation of the Rhodopsin-Transducin Interaction by a Highly Conserved Carboxylic Acid Group.
Biochemistry 32:7229–7236 (1993).
PMID: 8343512 [PubMed - indexed for MEDLINE]
24).
Min, K. C., Zyvaga, T. A., Cypess, A. M. & Sakmar, T. P.
Characterization of Mutant Rhodopsins Responsible for Autosomal Dominant Retinitis Pigmentosa. Mutations on the Cytoplasmic Surface Affect Transducin Activation.
J Biol Chem 268:9400–9404 (1993).
PMID: 8486634 [PubMed - indexed for MEDLINE]
23).
Zyvaga, T. A., Min, K. C., Beck, M. & Sakmar, T. P.
Movement of the Retinylidene Schiff Base Counterion in Rhodopsin by One Helix Turn Reverses the pH Dependency of the Metarhodopsin I to Metarhodopsin II Transition.
J Biol Chem 268:4661–4667 (1993). Correction: J Biol Chem 269:13056 (1994).
PMID: 8444840 [PubMed - indexed for MEDLINE]
22).
 Sakmar, T. P., Fahmy, K., Chan, T. & Lee, M.
Sakmar, T. P., Fahmy, K., Chan, T. & Lee, M.
Mutagenesis Studies of Rhodopsin Phototransduction.
in Structures and Functions of Retinal Proteins (Rigaud, J. L., ed.) Libbey, London, pp. 67-70 (1992).
[not indexed for MEDLINE]
21).

Sakmar, T. P.
The Traveler's Medical Kit.
Infect Dis Clin North Am 6:355–370 (1992).
PMID: 1624781 [PubMed - indexed for MEDLINE]
20).
Chan, T., Lee, M. & Sakmar, T. P.
Introduction of Hydroxyl-Bearing Amino Acids Causes Bathochromic Spectral Shifts in Rhodopsin: Amino Acid Substitutions Responsible for Red-Green Color Pigment Spectral Tuning.
J Biol Chem 267:9478–9480 (1992).
PMID: 1577792 [PubMed - indexed for MEDLINE]
19).
Sakmar, T. P., Franke, R. R. & Khorana, H. G.
Mutagenesis Studies of Rhodopsin Phototransduction.
in Signal Transduction in Photoreceptor Cells (Hargrave, P. A., Hofmann, K. P. & Kaupp, U. B., eds.) Springer-Verlag, Berlin, pp. 21–30 (1992).
[not indexed for MEDLINE]
18).
Lin, S. W., Sakmar, T. P., Franke, R. R., Khorana, H. G. & Mathies, R. A.
Resonance Raman Microprobe Spectroscopy of Rhodopsin Mutants: Effects of Substitutions in the Third Transmembrane Helix.
Biochemistry 31:5105–5111 (1992).
[not indexed for MEDLINE]
17).
Franke, R. R., Sakmar, T. P., Graham, R. M. & Khorana, H. G.
Structure and Function in Rhodopsin. Studies of the Interaction Between the Rhodopsin Cytoplasmic Domain and Transducin.
J Biol Chem 267:14767–14774 (1992).
PMID: 1634520 [PubMed - indexed for MEDLINE]
16).
Sakmar, T. P., Franke, R. R. & Khorana, H. G.
The Role of the Retinylidene Schiff Base Counterion in Rhodopsin in Determining Wavelength Absorbance and Schiff Base pKa.
Proc Natl Acad Sci U S A 88:3079–3083 (1991).
[not indexed for MEDLINE]
15).
Franke, R. R., König, B., Sakmar, T. P., Khorana, H. G. & Hofmann, K. P.
Rhodopsin Mutants That Bind But Fail to Activate Transducin.
Science 250:123–125 (1990).
PMID: 2218504 [PubMed - indexed for MEDLINE]
14).
Sakmar, T. P., Franke, R. R. & Khorana, H. G.
Glutamic Acid 113 Serves as the Retinylidene Schiff Base Counterion in Bovine Rhodopsin.
Proc Natl Acad Sci U S A 86:8309–8313 (1989).
PMID: 2573063 [PubMed - indexed for MEDLINE]
PMCID: PMC298270
13).
Karnik, S. S., Sakmar, T. P., Chen, H. -B. & Khorana, H. G.
Cysteine Residues 110 and 187 are Required for the Formation of Correct Structure in Bovine Rhodopsin.
Proc Natl Acad Sci U S A 85:8459–8463 (1988).
MID: 3186735 [PubMed - indexed for MEDLINE]
PMCID: PMC282477
12).
Sakmar, T. P. & Khorana, H. G.
Total Synthesis and Expression of a Gene for the a-Subunit of Bovine Rod Outer Segment Guanine Nucleotide-Binding Protein (Transducin).
Nucleic Acids Res 16:6361–6372 (1988).
[not indexed for MEDLINE]
11).
Franke, R. R., Sakmar, T. P., Oprian, D. D. & Khorana, H. G.
A Single Amino Acid Substitution in Rhodopsin ( Lysine 248 –-> Leucine) Prevents Activation of Transducin.
J Biol Chem 263:2119–2122 (1988).
PMID: 3123487 [PubMed - indexed for MEDLINE]
10).
Braiman, M., Bubis, J., Doi, T., Chen, H. -B., Flitsch, S. L., Franke, R. R., Gilles-Gonzalez, M. A., Graham, R. M., Karnik, S. S., Khorana, H. G., Knox, B. E., Krebs, M. P., Marti, T., Mogi, T., Nakayama, T., Oprian, D. D., Puckett, K. L., Sakmar, T. P., Stern, L. J., Subramaniam, S. & Thompson, D. A.
Studies on Light Transduction by Bacteriorhodopsin and Rhodopsin.
Cold Spring Harbor Symp Quant Biol 53:355–364 (1988).
[not indexed for MEDLINE]
9).
Franke, R. R., Sakmar, T. P., Graham, R. M. & Khorana, H. G.
Structure-Function Studies of Bovine Rhodopsin: Interactions with Transducin.
in Molecular Biology of the Eye: Genes, Vision and Ocular Disease(Piatigorsky, J., Zelenka, P. & Shinohara, T., eds.), Alan R. Liss, Inc., New York, pp. 45–52 (1988).
[not indexed for MEDLINE]
8).
Karnik, S. S., Sakmar, T. P., Franke, R. R., Knox, B. E., Oprian, D. D., Chen, H. -B. & Khorana, H. G.
Site-Specific Mutagenesis of Bovine Rhodopsin.
in Molecular Biology of the Eye: Genes, Vision and Ocular Disease (Piatigorsky, J., Zelenka, P., & Shinohara, T., eds.), Alan R. Liss, Inc., New York, pp. 23–34 (1988).
[not indexed for MEDLINE]
7).
Oprian, D., Sakmar, T. P., Nakayama, T., Chen, H. -B., Knox, B., Karnik, S., Franke, R. & Khorana, H. G.
Molecular Biological Studies of the Visual Pigment, Rhodopsin.
Discuss Neurosci 4:20–28 (1987).
[not indexed for MEDLINE]
6).
Ackerman, R. H., Richardson, E. P., Davis, K. R., Boom, W. H., Sakmar, T. P. & Haley, E. C.
Abrupt Onset of Headache Followed by Rapidly Progressive Encephalopathy in a 30-Year Old Woman - Angiitis of Central Nervous System (CPC).
N Engl J Med 313:566–575 (1985).
[not indexed for MEDLINE]
5).
Seid, R. C., Jr. & Sakmar, T. P.
A Differential Labeling Model for Determining the Number of Catalytically Essential Carboxyl Groups in Fumarase.
Biochim Biophys Acta (BBA) Enzymology 662:196–201 (1981).
[not indexed for MEDLINE]
4).
Randolph, A., Sakmar, T. P. & Heinrikson, R. L.
Phospholipases A2 - Structure, Function, and Evolution.
in Frontiers in Protein Chemistry (Liu, T. -Y. & Yasunobu, K., eds.) Elsevier, Amsterdam, pp. 297–322 (1980).
[not indexed for MEDLINE]
3).
Liang, S. M., Sakmar, T. P. & Liu, T. -Y.
Purification of an Endotoxin-Binding Protein from Limulus Amoebocyte Membranes.
J Biol Chem 255:5586–5590 (1980).
[not indexed for MEDLINE]
2).
Liu, T. -Y., Seid, R. C., Tai, J. Y., Liang, S. -M., Sakmar, T. P. & Robbins, J. B.
Studies on Limulus Lysate Coagulating System.
in Biomedical Applications of the Horseshoe Crab (Limulidae) (Cohen, E. B., ed.) Alan R. Liss, Inc., New York, pp. 147–158 (1979).
[not indexed for MEDLINE]
1).
Liu, T. -Y., Seid, R. C., Tai, J. Y., Liang, S. -M., Sakmar, T. P. & Robbins, J. B.
Studies on Limulus Lysate Coagulating System.
Prog Clin Biol Res 29:147–158 (1979).
[not indexed for MEDLINE]
3).
 Sakmar, T. P., Gardner, P. & Peterson, G. N.
Sakmar, T. P., Gardner, P. & Peterson, G. N.
Health Guide for International Travelers. 2nd edition.
Passport Books, National Textbook Co., Lincolnwood, IL. 171 pp. (1994).
2).

Sakmar, T. P., Gardner, P. & Peterson, G. N.
Health Guide for International Travelers.
Passport Books, National Textbook Co., Lincolnwood, IL. 143 pp. (1991 reprint).
1).
 Sakmar, T. P., Gardner, P. & Peterson, G. N.
Sakmar, T. P., Gardner, P. & Peterson, G. N.
Health Guide for International Travelers.
Passport Books, National Textbook Co., Lincolnwood, IL. 143 pp. (1989).
|

 Thomas Sakmar, M.D.
Thomas Sakmar, M.D. Marguerite Mangin, Ph.D.
Marguerite Mangin, Ph.D. Manija Kazmi, M.S.
Manija Kazmi, M.S. Thomas Huber, M.D., Ph.D.
Thomas Huber, M.D., Ph.D. Thomas Haines, Ph.D.
Thomas Haines, Ph.D. Pallavi Sachdev, Ph.D.
Pallavi Sachdev, Ph.D. Ruchi Gupta, Ph.D.
Ruchi Gupta, Ph.D. Saranga Naganathan, Ph.D.
Saranga Naganathan, Ph.D. Parag Mukhopadhyay, Ph.D.
Parag Mukhopadhyay, Ph.D. Neeraj Kapoor
Neeraj Kapoor Amy Grunbeck
Amy Grunbeck Adam Knepp
Adam Knepp Amy Ying Lin
Amy Ying Lin Thomas Haines
Thomas Haines Thomas Huber,
Thomas Huber,
 Neeraj Kapoor,
Neeraj Kapoor,
 Manija Kazmi
Manija Kazmi


















 Amy Ying Lin
Amy Ying Lin Pallavi Sachdev
Pallavi Sachdev
 Miller, J., Agarwal, A., Devi, L. A., Fontanini, K., Hamilton, J. A., Pin, J. P., Shields, D. C., Spek, C. A.,
Miller, J., Agarwal, A., Devi, L. A., Fontanini, K., Hamilton, J. A., Pin, J. P., Shields, D. C., Spek, C. A., Ahuja, S., Eilers, M., Hirshfeld, A.,
Ahuja, S., Eilers, M., Hirshfeld, A., 

 Ahuja, S., Hornak, V.,
Ahuja, S., Hornak, V.,  141).
141).

 Veldkamp, C. T.,
Veldkamp, C. T.,  Vogel, R., Mahalingam, M., Lüdeke, S.,
Vogel, R., Mahalingam, M., Lüdeke, S., 
 Louis, M.,
Louis, M., 


 Periole, X.,
Periole, X., 

 Vogel, R.,
Vogel, R.,  Botelho, A. V.,
Botelho, A. V., 
 Vogel, R., Siebert, F.,
Vogel, R., Siebert, F.,  Veldkamp, C. T.,
Veldkamp, C. T.,  Lewis, J. W., Szundi, I.,
Lewis, J. W., Szundi, I., 


 Vogel, R., Ludeke, S., Siebert, F.,
Vogel, R., Ludeke, S., Siebert, F., 

 Bartfai, T., Benovic, J. L., Bockaert, J., Bond, R. A., Bouvier, M., Christopoulos, A., Civelli, O., Devi, L. A., George, S. R., Inui, A., Kobilka, B., Leurs, R., Neubig, R., Pin, J. –P., Quirion, R., Roques, B. P.,
Bartfai, T., Benovic, J. L., Bockaert, J., Bond, R. A., Bouvier, M., Christopoulos, A., Civelli, O., Devi, L. A., George, S. R., Inui, A., Kobilka, B., Leurs, R., Neubig, R., Pin, J. –P., Quirion, R., Roques, B. P., 
 Billick, E.,
Billick, E., 



 Tsamis, F., Gavrilov, S., Kajumo, F.,
Tsamis, F., Gavrilov, S., Kajumo, F.,















 Isele, J.,
Isele, J.,  Unson, C. G., Wu, C. –R.,
Unson, C. G., Wu, C. –R.,  Cormier, E. G., Persuh, M., Thompson, D. A. D.,
Cormier, E. G., Persuh, M., Thompson, D. A. D.,  Ernst, O. E., Meyer, C.,
Ernst, O. E., Meyer, C., 



 Kochendoerfer, G. G.,
Kochendoerfer, G. G.,  Simpson, M. M., Ballesteros, J. A., Chiappa, V., Chen, J., Suehiro, M., Hartman, D. S., Godel, T.,
Simpson, M. M., Ballesteros, J. A., Chiappa, V., Chen, J., Suehiro, M., Hartman, D. S., Godel, T.,
 Beck, M., Siebert, F. &
Beck, M., Siebert, F. & 




 Garcia, P. D., Onrust, R., Bell, S. M.,
Garcia, P. D., Onrust, R., Bell, S. M., 






